Proteomic Analyses of Human Sperm Cells: Understanding the Role of Proteins and Molecular Pathways Affecting Male Reproductive Health
Ashok Agarwal, Manesh Kumar Panner Selvam and Saradha Baskaran, 27.02.2020
Abstract
Human sperm proteomics research has gained increasing attention lately, which provides complete information about the functional state of the spermatozoa. Changes in the sperm proteome are evident in several male infertility associated conditions. Global proteomic tools, such as liquid chromatography tandem mass spectrometry and matrix-assisted laser desorption/ionization time-of-flight, are used to profile the sperm proteins to identify the molecular pathways that are defective in infertile men. This review discusses the use of proteomic techniques to analyze the spermatozoa proteome. It also highlights the general steps involved in global proteomic approaches including bioinformatic analysis of the sperm proteomic data. Also, we have presented the findings of major proteomic studies and possible biomarkers in the diagnosis and therapeutics of male infertility. Extensive research on sperm proteome will help in understanding the role of fertility associated sperm proteins. Validation of the sperm proteins as biomarkers in different male infertility conditions may aid the physician in better clinical management.
AGARWAL, Ashok; PANNER SELVAM, Manesh Kumar; BASKARAN, Saradha. Proteomic analyses of human sperm cells: understanding the role of proteins and molecular pathways affecting male reproductive health. International journal of molecular sciences, 2020, 21. Jg., Nr. 5, S. 1621.
Publication: https://doi.org/10.3390/ijms21051621
 Disclaimer
Disclaimer
The publication Proteomic Analyses of Human Sperm Cells: Understanding the Role of Proteins and Molecular Pathways Affecting Male Reproductive Health by Ashok Agarwal, Manesh Kumar Panner Selvam and Saradha Baskaran is published under an open access license: https://creativecommons.org/licenses/by/4.0/. Granted rights: share — copy and redistribute the material in any medium or format and adapt — remix, transform, and build upon the material for any purpose, even commercially.
Curation by the MFGA team Relevant data sets presented in the publication have been identified. If possible, annotations (title, general information, conditions, processed tissue types and processed cell types) have been added based on information from the publication. Data tables and images that provide a good overview on the publication's findings on the data set have been extracted from the publication and/or supplement. If not stated otherwise, images are depicted with title and description exactly as in the publication. Tables have been adjusted to the MFGA table format. Conducted adjustments are explained in the detailed view of the tables. However, titles and descriptions have been adopted from the publication.
Data set 1:
Proteome: Other
Species
| Species |
|---|
| Human |
Cell Types
| Cell ontology | Maturity | Description | Species | Replicates | Cells per replicate |
|---|---|---|---|---|---|
| CL_0000019: sperm | Adult | human |
Images
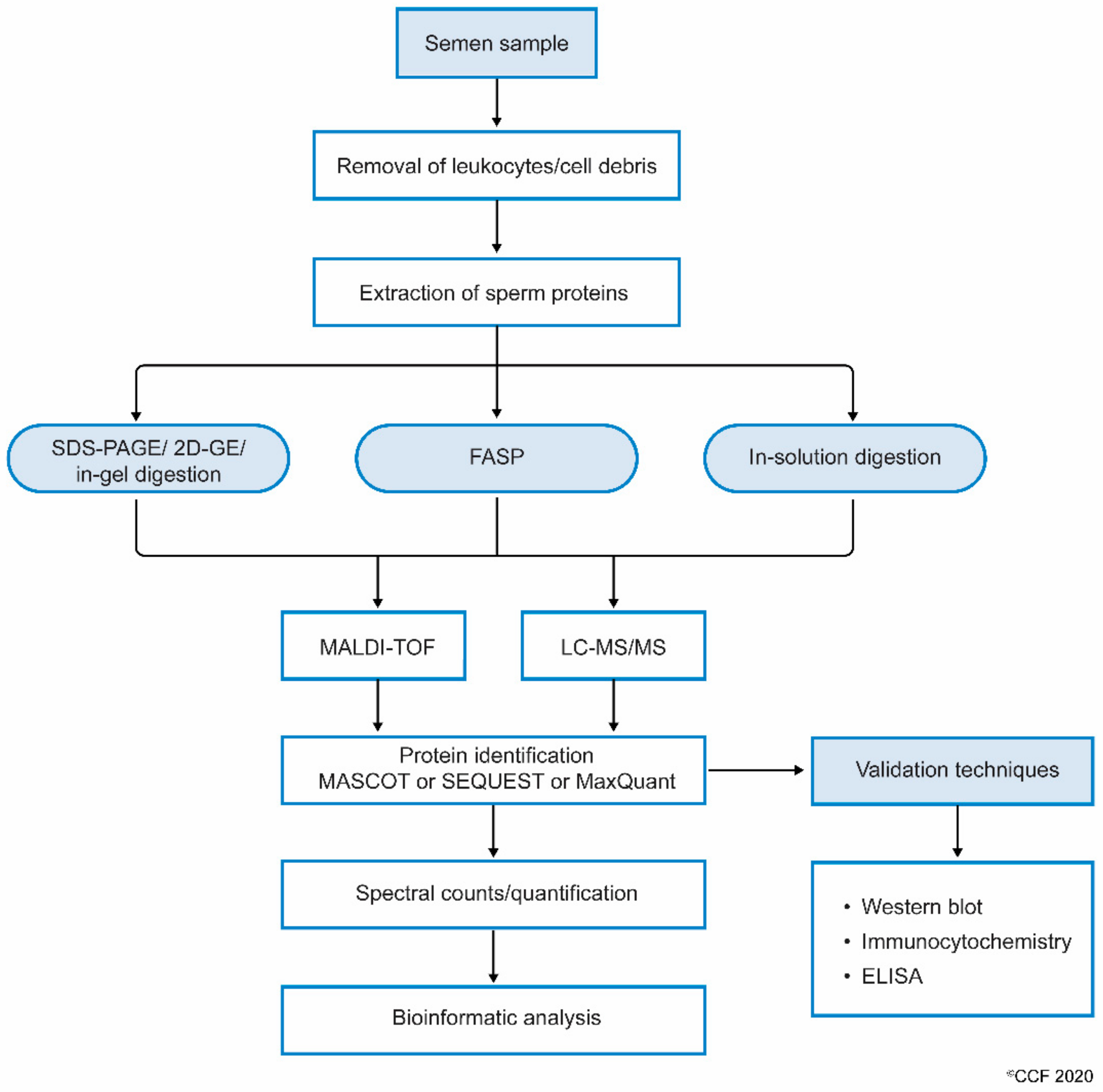
Figure 1: Workflow involving the processing of semen samples for sperm proteomics.
Licensed under: https://creativecommons.org/licenses/by/4.0/
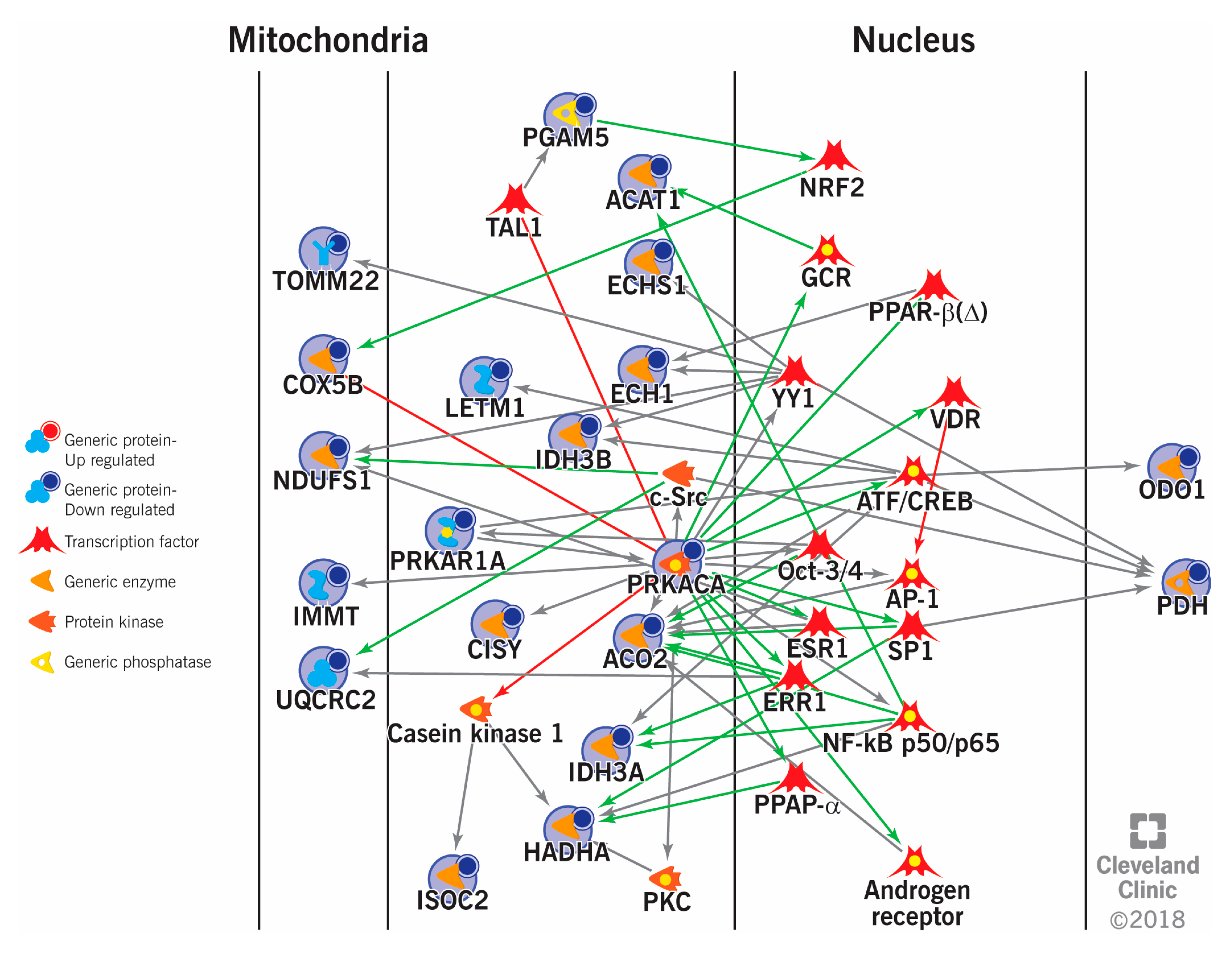
Figure 2: Interaction between the differentially expressed proteins and transcriptional factors in varicocele patients with mitochondrial dysfunction.
TOMM22: mitochondrial import receptor subunit TOM22 homolog, COX5B: cytochrome c oxidase subunit 5B, mitochondrial, NDUFS1: NADH-ubiquinone oxidoreductase 75 kDa subunit, mitochondrial, IMMT: NADH-ubiquinone oxidoreductase 75 kDa subunit, mitochondrial, UQCRC2: cytochrome b-c1 complex subunit 2, mitochondrial, PGAM5: serine/threonine-protein phosphatase PGAM5, mitochondrial, TAL1: T-cell acute lymphocytic leukemia protein 1, ACAT1: acetyl-CoA acetyltransferase, mitochondrial, ECHS1: enoyl-CoA hydratase, mitochondrial, ECH1: delta(3,5)-delta(2,4)-dienoyl-CoA isomerase, mitochondrial, LETM1: mitochondrial proton/calcium exchanger protein, IDH3B: isocitrate dehydrogenase [NAD] subunit beta, mitochondrial, c-Src: proto-oncogene tyrosine-protein kinase Src, PRKAR1A: cAMP-dependent protein kinase type I-alpha regulatory subunit, PRKACA: cAMP-dependent protein kinase catalytic subunit alpha, CISY: citrate synthase, mitochondrial, ACO2: aconitate hydratase, mitochondrial, IDH3A: isocitrate dehydrogenase [NAD] subunit alpha, mitochondrial, HADHA: trifunctional enzyme subunit alpha, mitochondrial, ISOC2: isochorismatase domain-containing protein 2, PKC: protein kinase C, NRF2: nuclear factor erythroid 2 (NFE2)-related factor 2, GCR: glucocorticoid receptor, PPAR-β (Δ): peroxisome proliferator-activated receptor beta or delta, YY1: Yin Yang 1, VDR: vitamin D receptor, ATF: activating transcription factor, CREB: cAMP response element binding, OCT-3/4: octamer-binding transcription factor 3/4, AP-1: activator protein 1, SP1: specificity protein 1, ESR1: estrogen receptor 1, ERR1: steroid hormone receptor ERR1, NF-κB: nuclear factor kappa-light-chain-enhancer of activated B cells.
Licensed under: https://creativecommons.org/licenses/by/4.0/
